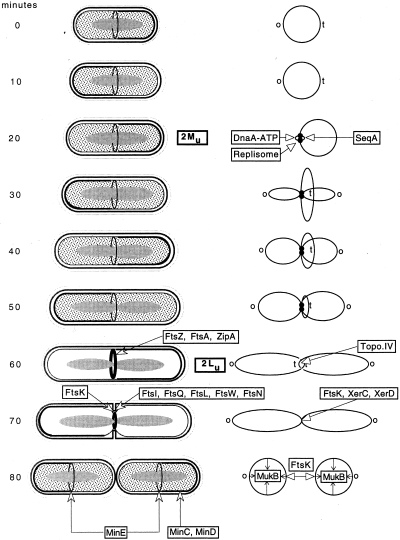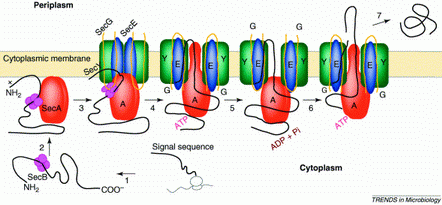
Fig. 1 the general secretion machinery in E.coli
(from Trends in Microbiology 9(10), p494)
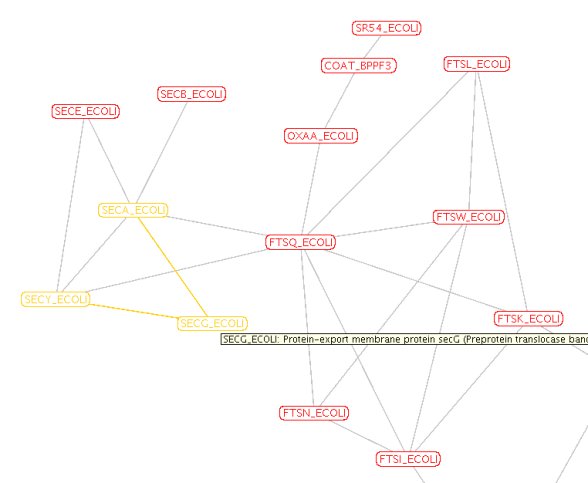
SecG, SecE and SecY build the
minimum set for translocation with SecA as the motor using ATP as fuel.
Additional Sec proteins may contribute for different kinds of
substrates, e.g.SecD.
The preprotein is represented
by a black line, with the gray region showing the signal sequence.
Steps 1-3, targeting. SecB, the Sec-system-specific chaperone, passes
the preprotein to the SecA and possibly already to the translocation
channel. SecA then binds to the membrane. SecY, SecE and SecG form the
transporter complex. Now, the preprotein gets fiddled through the
membrane like a thread through a needle.
Translocation is thought to occur in a step-wise fashion with a step of 20-30 amino acid residues at a time Therefore the whole process is repetitive. Little is known about the process, which occurs on the periplasmic side of the membrane (the inner membrane space), leading to the release and/or folding of the substrate protein.
Translocation is thought to occur in a step-wise fashion with a step of 20-30 amino acid residues at a time Therefore the whole process is repetitive. Little is known about the process, which occurs on the periplasmic side of the membrane (the inner membrane space), leading to the release and/or folding of the substrate protein.